CLC Genomics Workbench
Analyze high-throughput sequencing data.
$4995.00
CLC Genomics Workbench overview
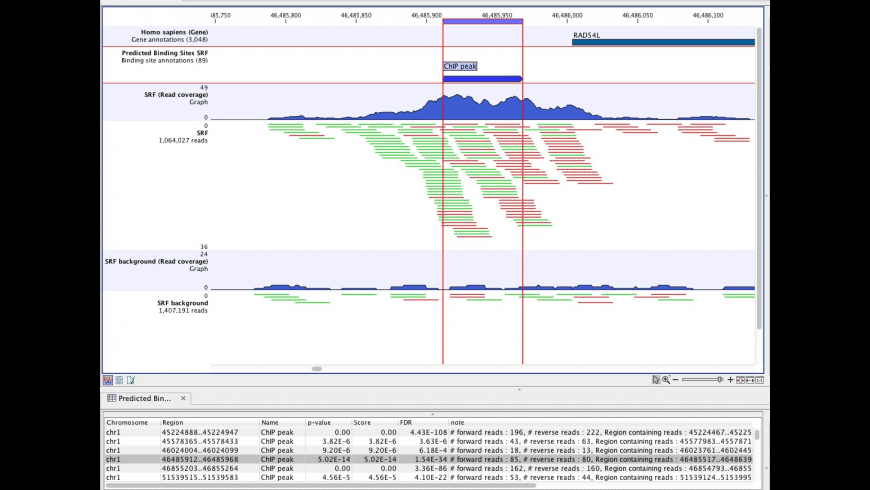
CLC Genomics Workbench incorporates cutting-edge technology and algorithms, while also supporting and integrating with the rest of your typical NGS workflow. CLC Genomics Workbench supports key next generation sequencing features within genomics, transcriptomics and epigenomics, and additionally it includes all the tools of CLC Main Workbench.
If you own a benchtop sequencing instrument you can make use of our 90-day trial license for CLC Genomics Workbench, please sign up at benchtopseq.com
What’s new in version 6.5.2
Updated on Jan 27 2014
Version 6.5.2: Bug fixes:
- Fixed: Error message when running analysis on experiments (statistical tests, clustering etc.)
- Fixed: track lists would sometimes be rendered empty when scrolling tracks with insertions.
- Fixed problem of unresponsive dialogs when running analysis with many reference sequences.
- Fixed a problem in track lists that caused the view to crash when there is an insertion at the very end of a chromosome.
- The folder used by the Workbench for storing log and settings files on Windows 8 has been updated to follow conventions for Windows 8 which is identical to Windows 7. Any existing settings will be copied to the new location automatically.
- Fixed various problems related to launching the Workbench through Java Webstart.
- Fixed: Opening a search view for searching sequences at NCBI would sometimes fail.
- Fixed: The Target Regions Statistics tool did not handle annotations covering the starting point of circular reference sequences properly. If you are using the tool with annotations spanning across the starting point of a circular reference, we recommend re-running the analysis.
- In BAM files created by BWA, non-specific reads are now recognized as such during import. Previously, they were treated as unique reads.
- Improved stability of Probabilistic variant detection on huge data sets.
- Fixed various stability and performance problems of Maximum likelihood phylogeny.
- Fixed problem that caused a crash with extract consensus sequence tool with certain parameter configurations and with read mappings with no reads.
- Fixed crash of detailed mapping report tool with certain data sets.
- Fixed error in importing SOLiD XSQ files.
- Fixed an error when importing BAM files, including problems regarding download of reference sequences.
- Fixed a read mapper error occurring under special circumstances when excluding regions of a reference when mapping reads .
- Fixed a bug in the Assemble Sequences tool causing some data sets to produce inferior contigs.
- This release is based on CLC Assembly Cell 4.2.1
- This release can be used with CLC Genomics Server 5.5.X.
Information
License
Demo
Size
190 MB
Developer’s website
http://www.clcbio.com/products/clc-genomics-workbench/Downloads
528
App requirements
- Intel 64
- Intel 32
- PPC 32
- Mac OS X 10.5.8 or later
Try our new feature and write a detailed review about CLC Genomics Workbench. All reviews will be posted soon.
(0 Reviews of )
There are no reviews yet
Comments
User Ratings
Help the community
There are no reviews yet, be the first to leave one
Help the community
There are no ratings yet, be the first to leave one
$4995.00
Similar apps
Be the first one to propose an app
similar to CLC Genomics Workbench.
similar to CLC Genomics Workbench.
New and Recently Updated
ndCurveMaster
Unique multivariable curve fitting program that automates the fitting of nonlinear. regression equations to your datasets, accommodating an unlimited number of input variables.